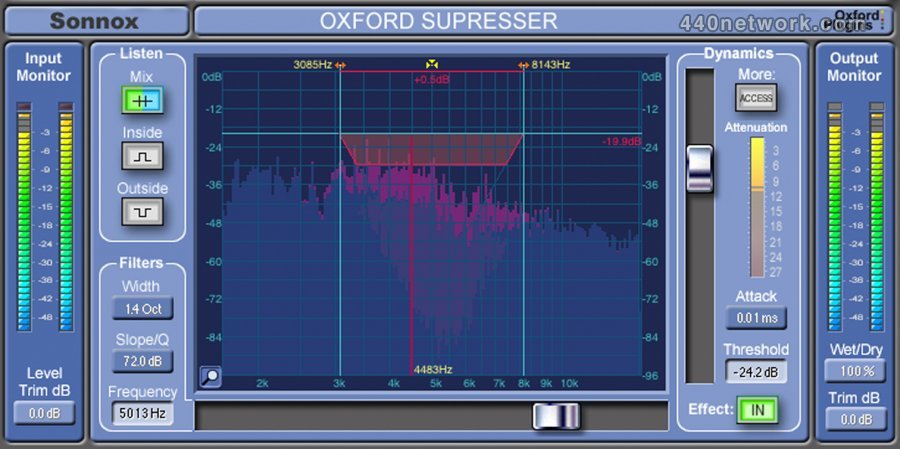 
We further use the 2′-hydroxyl reactive SHAPE reagent acetylimidazole to probe RNA structure at the single-molecule level with readout by direct nanopore sequencing. We detect endogenous 2′- O-methyl and base modifications across E. coli and S. cerevisiae ribosomal RNAs as shifts in current signal and dwell times distally through interactions with the helicase motor protein. We cannot confirm if there is a free download of this software available. Using the link below to download Sonnox Oxford R3 Dynamics Native VST from the developers website was possible when we last checked. 
Here, we demonstrate direct RNA nanopore sequencing to detect endogenous and exogenous RNA modifications on long RNAs at the single-molecule level. Download Sonnox Oxford R3 Dynamics Native VST Thank you for using our software library. Many methods exist to detect RNA modifications by short-read sequencing however, limitations on read length inherent to short-read-based methods dissociate modifications from their native context, preventing single-molecule modification analysis. In addition, modifications can be introduced to RNA molecules using chemical probing strategies that reveal RNA structure and dynamics. These modifications modulate diverse biological processes such as genetic recoding and mRNA export and folding. Modifications are present on many classes of RNA, including tRNA, rRNA, and mRNA.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
